Editor’s Note: when you’re in a resource-poor field situation—one where you’re trying to trace the spread of an epidemic or the impact a drug has on resistance to an epidemic—be able to do rapid genomic analyzes with equipment you can carry with you The backpack is extremely useful. As we begin to explore the diversity of life forms, the ability to have fast, autonomous devices (tricorders) to test in situ will be critical. How we’re already doing this on Earth can serve as an analogy for the things we’ll need to design as we equip missions that set out to explore new worlds.
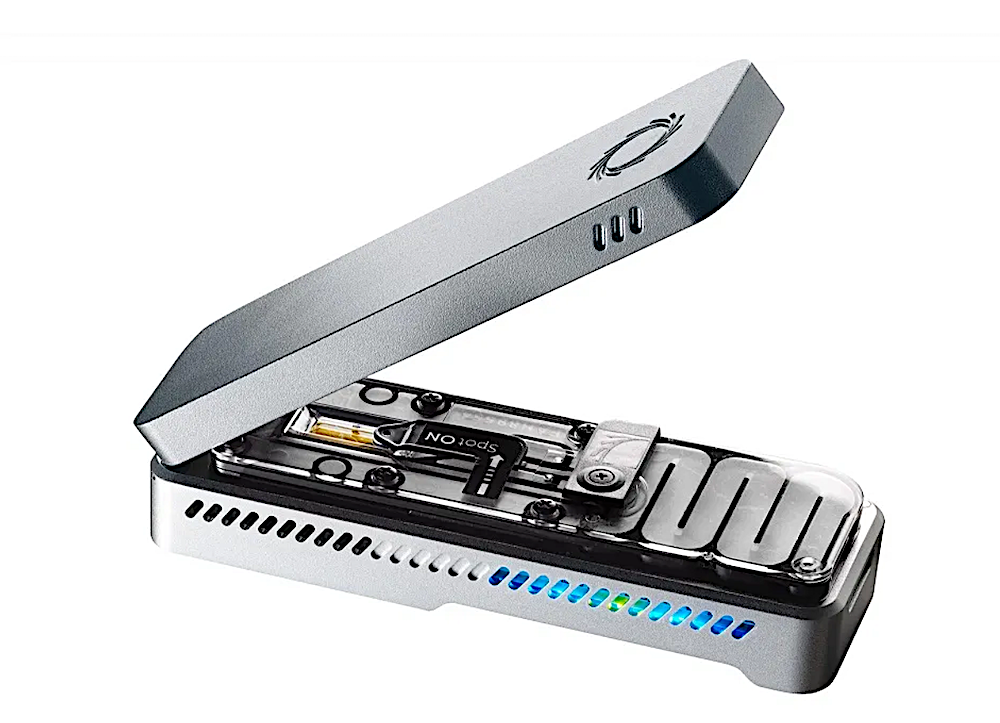
Scientists have developed a technique to rapidly and reliably detect genetic changes in malaria parasites in Ghana, using only a gaming laptop and portable MinION sequencer from Oxford Nanopore1.
Researchers from the Wellcome Sanger Institute and the University of Ghana have been able to demonstrate for the first time that end-to-end real-time pathogen monitoring from clinical blood samples is possible in resource-limited rural malaria hotspots. They looked for key markers of drug resistance in the malaria parasite and diversity in the vaccine’s target gene2.
The findings, published in Nature Microbiology (Nov. 23), pave the way for local monitoring of drug resistance and evaluation of new malaria vaccines in affected areas.
Despite intensive control efforts in recent decades, malaria still kills over 600,000 people annually3, most of whom are young children in sub-Saharan Africa. A key factor is the ability of malaria parasites to quickly develop resistance to antimalarial drugs and other medical interventions.
Genomic surveillance – the continuous monitoring of changes in the parasite’s DNA – provides the tools to analyze the genomic data behind parasite resistance. But until recently, it was mostly done in laboratories in high-income, non-malaria-endemic countries, concentrating capacity away from affected areas.
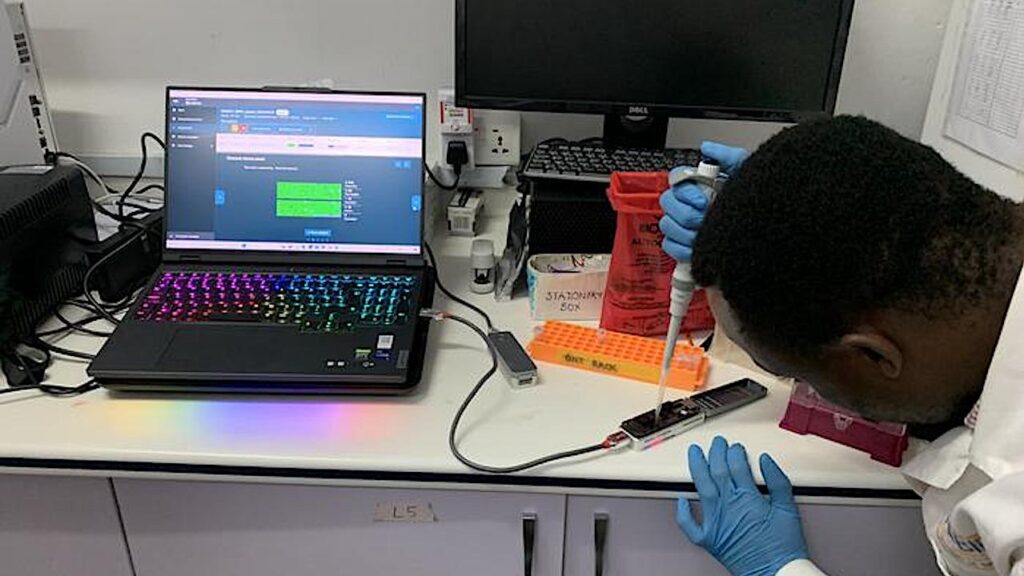

Edem Adika loads a flow cell in Accra CREDIT Dr William Hamilton
In this new study, researchers set out to develop an accessible near-real-time technology to track parasite mutations in the communities most affected by malaria.
The researchers used standard molecular biology equipment to collect parasites from blood spot samples, prepared with a simple finger prick. They then sequenced and analyzed the DNA of the malaria parasite using the MinION handheld device and a laptop to detect known markers of drug resistance, emerging mutations and targets for new malaria vaccines.
The team successfully conducted the study from two locations: an urban hospital in Ghana’s capital Accra and a rural town 11 hours’ drive to the north. They were able to generate sequence information in just 48 hours after receiving a sample, while keeping costs to a minimum, at around £27 per sample in batches of 96.
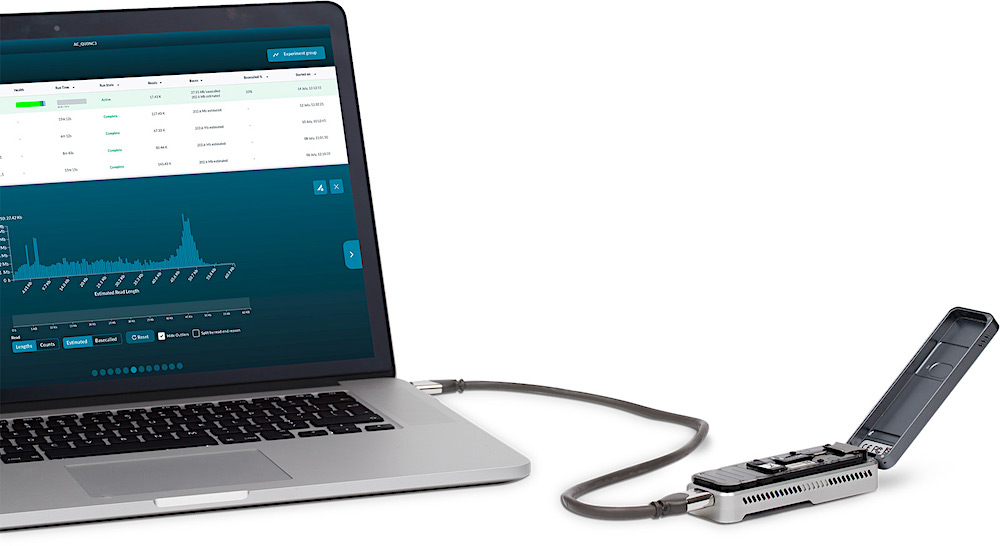

MinION DNA Sequencer More technical information
The team showed that first-line treatments remain broadly effective against local strains of malaria in Ghana currently4. However, continued monitoring is necessary, including protection of high-risk groups receiving targeted interventions5.
They also discovered multiple genetic differences between circulating malaria strains and the protein targeted by newly recommended malaria vaccines. Although they require further study, these could affect the latest vaccine rollout across Africa.
Edem Adika, co-first author of the study at the University of Ghana, said: “By taking the sequence at the source, the information arrives in days rather than years – enabling rapid, local responses. This unprecedented speed promises to be a powerful game-changer against infectious diseases that outsmart our countermeasures. We hope that this in-situ approach will soon be applied here to other pathogens.”
Dr William Hamilton, senior author of the study at the Wellcome Sanger Institute, said: “The repeated evolution and spread of resistance to key antimalarial drugs has hampered efforts to eradicate malaria over the past 70 years. Expanding molecular surveillance in Africa is now critical to monitor emerging resistance to drugs and diagnostic tests and to inform interventions such as new vaccines.”
Dr Lucas Amenga-Etego, senior author of the study at the West African Center for Cell Biology of Infectious Pathogens (WACCBIP), University of Ghana, said: “This sequencing workflow has enormous potential to address the sequencing gap in sub-Saharan Africa given its lower cost and ease of use. However, sustainable capacity scaling requires expanding local training programs, bioinformatics infrastructure, and data science expertise. These should be priorities for the global pathogen genomics community going forward.”
Notes to editors:
- The Oxford Nanopore MinION is a portable, pocket-sized DNA sequencer that uses nanopore sequencing technology to analyze genetic material. This technology offers advantages such as long-term sequencing reads and immediate detection of genetic changes.
- The assay designed for this procedure screened parasite samples using a polymerase chain reaction (PCR), looking at five key markers in DNA associated with changes that conferred drug resistance, as well as the primary vaccine and monoclonal antibody target, the circumsporozoite protein (csp). csp helps parasites establish infection early after injection by feeding on mosquitoes, so blocking csp helps prevent malaria infections.
- [74n5c4m7.r.eu-west-1.awstrack.me]
- The team found no evidence of resistance to artemisinins – the current first-line and best treatment for P. falciparum. Partial resistance to artemisinins has been detected in southeast Asia and eastern Africa. Continued monitoring for resistance will be necessary.
- Although mutations associated with resistance to the antimalarial drugs sulfadoxine and pyrimethamine (SP) were found, the particularly dangerous mutations that cause “high-level” resistance to SP were not found. This is reassuring because SP is used for prophylaxis in young children and during pregnancy in Ghana.
Publication:
ST Girgis et al. (2023)’Drug resistance and vaccine target surveillance of Plasmodium falciparum using nanopore sequencing in Ghana.’ Nature Microbiology. DOI: 10.1038/s41564-023-01516-6. (open access)
Financing:
This research was supported by Wellcome and a grant for Human Heredity and Health in Africa (H3Africa). For full funding acknowledgement, see publication.
Astrobiology, Genomics, Epidemiology,


